RIXS $L_{2,3}M_{4,5}$
Using the function ResonantSpectra we can calculate inelastic x-ray scattering. (or other second order processess) Here we show the example of $d$-$d$ excitations in NiO using RIXS. Due to an ever increasing resolution this method gains rapidly in popularity and impact. See for example the combined work of Ghiringhelli \textit{et al.} For NiO see \cite{Ghiringhelli:2005kp}.
A script to calculate these spectra is:
- RIXS_L23M45.Quanty
-- this example calculates the resonant inelastic x-ray scattering in NiO we look at the -- Ni L23M45 edge, i.e. we make an excitation from 2p to 3d (L23) and decay from the -- 3d shell back to the 2p shell (final "hole" in the 3d shell M45 edge). These spectra -- measure d-d excitations and or magnons. -- we minimize the output by setting the verbosity to 0 Verbosity(0) -- in order to do this calculation we need a Ni 2p shell and a Ni 3d shell NF=16 NB=0 IndexDn_2p={0,2,4} IndexUp_2p={1,3,5} IndexDn_3d={6,8,10,12,14} IndexUp_3d={7,9,11,13,15} -- just like in the previous example we define several operators acting on the Ni -3d shell OppSx =NewOperator("Sx" ,NF, IndexUp_3d, IndexDn_3d) OppSy =NewOperator("Sy" ,NF, IndexUp_3d, IndexDn_3d) OppSz =NewOperator("Sz" ,NF, IndexUp_3d, IndexDn_3d) OppSsqr =NewOperator("Ssqr" ,NF, IndexUp_3d, IndexDn_3d) OppSplus=NewOperator("Splus",NF, IndexUp_3d, IndexDn_3d) OppSmin =NewOperator("Smin" ,NF, IndexUp_3d, IndexDn_3d) OppLx =NewOperator("Lx" ,NF, IndexUp_3d, IndexDn_3d) OppLy =NewOperator("Ly" ,NF, IndexUp_3d, IndexDn_3d) OppLz =NewOperator("Lz" ,NF, IndexUp_3d, IndexDn_3d) OppLsqr =NewOperator("Lsqr" ,NF, IndexUp_3d, IndexDn_3d) OppLplus=NewOperator("Lplus",NF, IndexUp_3d, IndexDn_3d) OppLmin =NewOperator("Lmin" ,NF, IndexUp_3d, IndexDn_3d) OppJx =NewOperator("Jx" ,NF, IndexUp_3d, IndexDn_3d) OppJy =NewOperator("Jy" ,NF, IndexUp_3d, IndexDn_3d) OppJz =NewOperator("Jz" ,NF, IndexUp_3d, IndexDn_3d) OppJsqr =NewOperator("Jsqr" ,NF, IndexUp_3d, IndexDn_3d) OppJplus=NewOperator("Jplus",NF, IndexUp_3d, IndexDn_3d) OppJmin =NewOperator("Jmin" ,NF, IndexUp_3d, IndexDn_3d) Oppldots=NewOperator("ldots",NF, IndexUp_3d, IndexDn_3d) -- as in the previous example we define the Coulomb interaction OppF0 =NewOperator("U", NF, IndexUp_3d, IndexDn_3d, {1,0,0}) OppF2 =NewOperator("U", NF, IndexUp_3d, IndexDn_3d, {0,1,0}) OppF4 =NewOperator("U", NF, IndexUp_3d, IndexDn_3d, {0,0,1}) -- as in the previous example we define the crystal-field operator Akm = PotentialExpandedOnClm("Oh", 2, {0.6,-0.4}) OpptenDq = NewOperator("CF", NF, IndexUp_3d, IndexDn_3d, Akm) -- and as in the previous example we define operators that count the number of eg and t2g -- electrons Akm = PotentialExpandedOnClm("Oh", 2, {1,0}) OppNeg = NewOperator("CF", NF, IndexUp_3d, IndexDn_3d, Akm) Akm = PotentialExpandedOnClm("Oh", 2, {0,1}) OppNt2g = NewOperator("CF", NF, IndexUp_3d, IndexDn_3d, Akm) -- In order te describe the resonance we need the interaction on the 2p shell (spin-orbit) Oppcldots= NewOperator("ldots", NF, IndexUp_2p, IndexDn_2p) -- and the Coulomb interaction between the 2p and 3d shell OppUpdF0 = NewOperator("U", NF, IndexUp_2p, IndexDn_2p, IndexUp_3d, IndexDn_3d, {1,0}, {0,0}) OppUpdF2 = NewOperator("U", NF, IndexUp_2p, IndexDn_2p, IndexUp_3d, IndexDn_3d, {0,1}, {0,0}) OppUpdG1 = NewOperator("U", NF, IndexUp_2p, IndexDn_2p, IndexUp_3d, IndexDn_3d, {0,0}, {1,0}) OppUpdG3 = NewOperator("U", NF, IndexUp_2p, IndexDn_2p, IndexUp_3d, IndexDn_3d, {0,0}, {0,1}) -- next we define the dipole operator. The dipole operator is given as epsilon.r -- with epsilon the polarization vector of the light and r the unit position vector -- We can expand the position vector on (renormalized) spherical harmonics and use -- the crystal-field operator to create the dipole operator. -- x polarized light is defined as x = Cos[phi]Sin[theta] = sqrt(1/2) ( C_1^{(-1)} - C_1^{(1)}) Akm = {{1,-1,sqrt(1/2)},{1, 1,-sqrt(1/2)}} TXASx = NewOperator("CF", NF, IndexUp_3d, IndexDn_3d, IndexUp_2p, IndexDn_2p, Akm) -- y polarized light is defined as y = Sin[phi]Sin[theta] = sqrt(1/2) I ( C_1^{(-1)} + C_1^{(1)}) Akm = {{1,-1,sqrt(1/2)*I},{1, 1,sqrt(1/2)*I}} TXASy = NewOperator("CF", NF, IndexUp_3d, IndexDn_3d, IndexUp_2p, IndexDn_2p, Akm) -- z polarized light is defined as z = Cos[theta] = C_1^{(0)} Akm = {{1,0,1}} TXASz = NewOperator("CF", NF, IndexUp_3d, IndexDn_3d, IndexUp_2p, IndexDn_2p, Akm) -- besides linear polarized light one can define circular polarized light as the sum of -- x and y polarizations with complex prefactors TXASr = sqrt(1/2)*(TXASx - I * TXASy) TXASl =-sqrt(1/2)*(TXASx + I * TXASy) -- we can remove zero's from the dipole operator by chopping it. TXASr.Chop() TXASl.Chop() -- the 3d to 2p dipole transition is the conjugate transpose of the 2p to 3d dipole transition TXASxdag = ConjugateTranspose(TXASx) TXASydag = ConjugateTranspose(TXASy) TXASzdag = ConjugateTranspose(TXASz) TXASldag = ConjugateTranspose(TXASl) TXASrdag = ConjugateTranspose(TXASr) -- once all operators are defined we can set some parameter values. -- the value of U drops out of a crystal-field calculation as the total number of electrons -- is always the same U = 0.000 -- F2 and F4 are often referred to in the literature as J_{Hund}. They represent the energy -- differences between different multiplets. Numerical values can be found in the back of -- my PhD. thesis for example. http://arxiv.org/abs/cond-mat/0505214 F2dd = 11.142 F4dd = 6.874 -- F0 is not the same as U, although they are related. Unimportant in crystal-field theory -- the difference between U and F0 is so important that I do include it here. (U=0 so F0 is not) F0dd = U+(F2dd+F4dd)*2/63 -- in crystal field theory U drops out of the equation, also true for the interaction between the -- Ni 2p and Ni 3d electrons Upd = 0.000 -- The Slater integrals between the 2p and 3d shell, again the numerical values can be found -- in the back of my PhD. thesis. (http://arxiv.org/abs/cond-mat/0505214) F2pd = 6.667 G1pd = 4.922 G3pd = 2.796 -- F0 is not the same as U, although they are related. Unimportant in crystal-field theory -- the difference between U and F0 is so important that I do include it here. (U=0 so F0 is not) F0pd = Upd + G1pd*1/15 + G3pd*3/70 -- tenDq in NiO is 1.1 eV as can be seen in optics or using IXS to measure d-d excitations tenDq = 1.100 -- the Ni 3d spin-orbit is small but finite zeta_3d = 0.081 -- the Ni 2p spin-orbit is very large and should not be scaled as theory is quite accurate here zeta_2p = 11.498 -- we can add a small magnetic field, just to get nice expectation values. (units in eV... ) Bz = 0.000001 -- the total Hamiltonian is the sum of the different operators multiplied with their prefactor Hamiltonian = F0dd*OppF0 + F2dd*OppF2 + F4dd*OppF4 + tenDq*OpptenDq + zeta_3d*Oppldots + Bz*(2*OppSz+OppLz) -- We normally do not include core-valence interactions between filed and partially filled -- shells as it drops out anyhow. For the XAS we thus need to define a "different" -- (more complete) Hamiltonian. XASHamiltonian = Hamiltonian + zeta_2p * Oppcldots + F0pd * OppUpdF0 + F2pd * OppUpdF2 + G1pd * OppUpdG1 + G3pd * OppUpdG3 -- We saw in the previous example that NiO has a ground-state doublet with occupation -- t2g^6 eg^2 and S=1 (S^2=S(S+1)=2). The next state is 1.1 eV higher in energy and thus -- unimportant for the ground-state upto temperatures of 10 000 Kelvin. We thus restrict -- the calculation to the lowest 3 eigenstates. Npsi=3 -- in order to make sure we have a filling of 8 -- electrons we need to define some restrictions -- We need to restrict the occupation of the Ni-2p shell to 6 for the ground state and for -- the Ni 3d-shell to 8. StartRestrictions = {NF, NB, {"111111 0000000000",6,6}, {"000000 1111111111",8,8}} -- And calculate the lowest 3 eigenfunctions psiList = Eigensystem(Hamiltonian, StartRestrictions, Npsi) -- In order to get some information on these eigenstates it is good to plot expectation values -- We first define a list of all the operators we would like to calculate the expectation value of oppList={Hamiltonian, OppSsqr, OppLsqr, OppJsqr, OppSz, OppLz, Oppldots, OppF2, OppF4, OppNeg, OppNt2g}; -- next we loop over all operators and all states and print the expectation value print(" <E> <S^2> <L^2> <J^2> <S_z> <L_z> <l.s> <F[2]> <F[4]> <Neg> <Nt2g>"); for i = 1,#psiList do for j = 1,#oppList do expectationvalue = Chop(psiList[i]*oppList[j]*psiList[i]) io.write(string.format("%8.3f ",Complex.Re(expectationvalue))) end io.write("\n") end -- Not the main task of this example, but in order to know where the resonance sits it is -- nice to calculate the x-ray absorption spectra XASSpectra = CreateSpectra(XASHamiltonian, TXASx, psiList[1], {{"Emin",-10}, {"Emax",20}, {"NE",3500}, {"Gamma",1.0}}) XASSpectra.Print({{"file", "RIXSL23M45_XAS.dat"}}) -- We also calculate the fluorescence yield spectra which are equal to the integral over the -- RIXS spectra FYSpectra = CreateFluorescenceYield(XASHamiltonian, TXASx, TXASydag, psiList[1], {{"Emin",-10}, {"Emax",20}, {"NE",3500}, {"Gamma",1.0}}) FYSpectra.Print({{"file","RIXSL23M45_FY.dat"}}) -- and we calculate the RIXS spectra. Note that in order to calculate RIXS you need to -- specify two Hamiltonians. -- These calculations can be lengthy and one should think that for each incoming energy -- (E1) one needs to a new calculation. In this case there are thus 120 calculations RIXSSpectra = CreateResonantSpectra(XASHamiltonian, Hamiltonian, TXASx, TXASydag, psiList[1], {{"Emin1",-10}, {"Emax1",20}, {"NE1",120}, {"Gamma1",1.0}, {"Emin2",-0.5}, {"Emax2",7.5}, {"NE2",8000}, {"Gamma2",0.05}}) RIXSSpectra.Print({{"file", "RIXSL23M45.dat"}}) print("Finished calculating the spectra now start plotting.\nThis might take more time than the calculation"); -- Once finished we can make some nice plots. The spectra are saved to disk in ASCII format -- but I like to add gnuplot scripts so you can look at pictures imeidately gnuplotInput = [[ set autoscale set xtic auto set ytic auto set style line 1 lt 1 lw 1 lc rgb "#000000" set style line 2 lt 1 lw 1 lc rgb "#FF0000" set style line 3 lt 1 lw 1 lc rgb "#00FF00" set style line 4 lt 1 lw 1 lc rgb "#0000FF" set out 'RIXSL23M45_Map.ps' set terminal postscript portrait enhanced color "Times" 8 size 7.5,6 unset colorbox energyshift=857.6 set multiplot set size 0.5,0.55 set origin 0,0 set ylabel "resonant energy (eV)" font "Times,10" set xlabel "energy loss (eV)" font "Times,10" set yrange [852:860] set xrange [-0.5:7.5] plot "<(awk '((NR>5)&&(NR<8007)){for(i=3;i<=NF;i=i+2){printf \"%s \", $i}{printf \"%s\", RS}}' RIXSL23M45.dat)" matrix using ($2/1000-0.5):($1/4+energyshift-10):(-$3) with image notitle set origin 0.5,0 set yrange [869:877] set xrange [-0.5:7.5] plot "<(awk '((NR>5)&&(NR<8007)){for(i=3;i<=NF;i=i+2){printf \"%s \", $i}{printf \"%s\", RS}}' RIXSL23M45.dat)" matrix using ($2/1000-0.5):($1/4+energyshift-10):(-$3) with image notitle unset multiplot set out 'RIXSL23M45_Spec.ps' set terminal postscript portrait enhanced color "Times" 8 size 7.5,5 set multiplot set size 0.25,1.0 set origin 0,0 set ylabel "E (eV)" font "Times,10" set xlabel "XAS" font "Times,10" set yrange [energyshift-10:energyshift+20] set xrange [-0.3:0] plot "RIXSL23M45_XAS.dat" using 3:($1+energyshift) notitle with lines ls 1,\ "RIXSL23M45_FY.dat" using (-3*$2):($1+energyshift) notitle with lines ls 4 set size 0.8,1.0 set origin 0.2,0.0 set xlabel "Energy loss (eV)" font "Times,10" unset ylabel unset ytics set xrange [-0.5:7.5] ofset = 0.25 scale=3 plot for [i=0:120] "RIXSL23M45.dat" using 1:(-column(3+2*i)*scale+ofset*i-10 + energyshift) notitle with lines ls 4 unset multiplot ]] -- write the gnuplot script to a file file = io.open("RIXSL23M45.gnuplot", "w") file:write(gnuplotInput) file:close() -- call gnuplot to execute the script os.execute("gnuplot RIXSL23M45.gnuplot") -- transform to pdf and eps os.execute("ps2pdf RIXSL23M45_Map.ps ; ps2eps RIXSL23M45_Map.ps ; mv RIXSL23M45_Map.eps temp.eps ; eps2eps temp.eps RIXSL23M45_Map.eps ; rm temp.eps") os.execute("ps2pdf RIXSL23M45_Spec.ps ; ps2eps RIXSL23M45_Spec.ps ; mv RIXSL23M45_Spec.eps temp.eps ; eps2eps temp.eps RIXSL23M45_Spec.eps ; rm temp.eps")
It produces two plots.
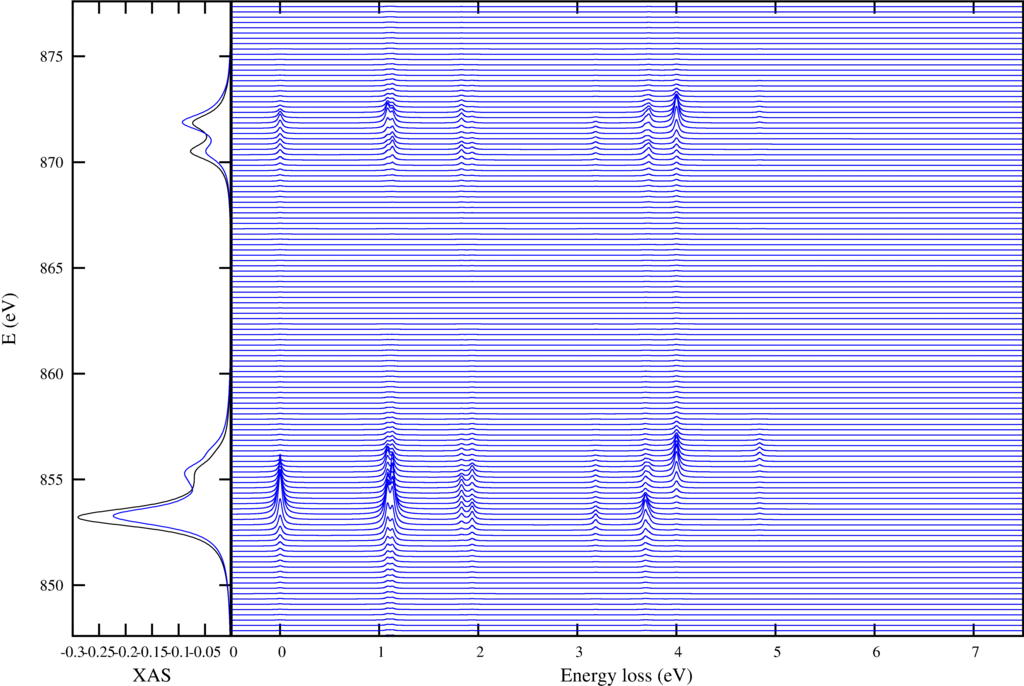
The first shows on the left the XAS spectra in black and the integrated RIXS spectra (FY) in blue. The resonant enhanced $d$-$d$ excitations are shown in the right plot.
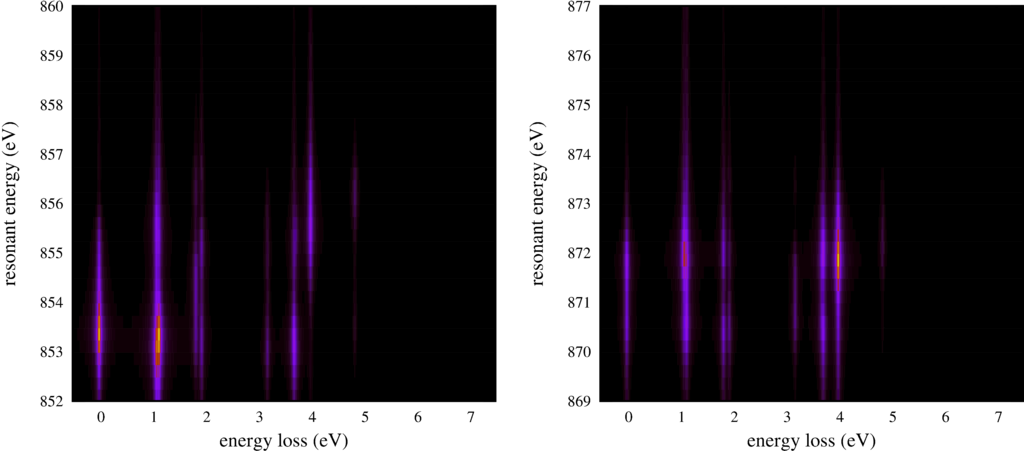
The second plot shows the same RIXS spectra, but now as an intensity map.
The text output of the script is:
- RIXS_L23M45.out
<E> <S^2> <L^2> <J^2> <S_z> <L_z> <l.s> <F[2]> <F[4]> <Neg> <Nt2g> -2.721 1.999 12.000 15.120 -0.994 -0.286 -0.324 -1.020 -0.878 2.011 5.989 -2.721 1.999 12.000 15.120 -0.000 -0.000 -0.324 -1.020 -0.878 2.011 5.989 -2.721 1.999 12.000 15.120 0.994 0.286 -0.324 -1.020 -0.878 2.011 5.989 Start of LanczosTriDiagonalizeKrylovRR Start of LanczosTriDiagonalizeKrylovRR Finished calculating the spectra now start plotting. This might take more time than the calculation